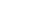
大规模蛋白质相互作用研究方法进展(四)
表1 蛋白质相互作用分析相关数据库及网站
网站 资源类型 网址
DIP 蛋白质相互作用http: //dip.doe-mbi.uda.edu
INTERACT 蛋白质相互作用http: //bioinf.man.ac.uk/interactpr.htm
ProNet 蛋白质相互作用http: //pronet.doubletwist.com/
MIPS 蛋白质相互作用http:
//www.mips.biochem.mpg.de/proj/yeast/tables/interactoin/index.htm
Proteome 蛋白质相互作用http: //www.proteome.com
Bind 蛋白质相互作用http: //www.binddb.org
String 基因共定位http: //www.bork.embl-heidelberg.de/string/
CoGs 种系发生谱http: //www.ncbi.nlm.nih.gov/CoG/
2.6 其他方法 具有相关功能或相互作用的蛋白质他们的基因有可能受到了相同的调控,在进化过程中也保持一定的相似性。因此,可以通过检测 mRNA
在特定时期的表达量,分析相关的基因产物是否具有相互作用关系。Stuart 等[33]研究了人、果蝇及线虫的基因共表达模式,发现22
163 种共表达关系并且在进化上它们也表现出一定的保守性。必须指出的是,有些蛋白的表达与mRNA可能不存在相互关系,mRNA 的共表达不反映蛋白间的相互作用;另外所得数据的分析也是一个繁琐的任务。
除了上述大规模分析蛋白质相互作用的方法外,最近还涌现出一些其他的方法。多维蛋白鉴定技术(multidimensional
protein identification technology,
MudPIT)[34],通过确定蛋白质的亚细胞定位来研究其相互作用等[35]。免疫共沉淀、GST
pull-down、 Far-western、化学交联等目前虽不适于大规模研究,但小范围的分析方法可对得到的结果进行补充印证。众多的研究方法可以从不同的角度研究蛋白质组中的蛋白相互作用,弥补单个方法的不足,从而得到尽可能完整的蛋白质相互作用图谱。
3 展望
高等动物基因组研究表明,调节生物体复杂性的不是基因数量的增加,而是基于更为复杂的蛋白质-蛋白质间发生的相互作用。建立起这些分子之间的相互作用及调节网络是后基因组时代的首要任务。这种相互作用的研究包括稳定复合物的亲和力、半衰期及瞬间的相互作用,而且这些复合物中蛋白质的相互作用不是一成不变的,他们存在着动态变化。因此,最终的目标是通过这些作用网络,加上基因动态表达研究、蛋白质表达量分析、蛋白定位以及基因功能分析等,最终了解细胞内各种生理反应的发生及调节机制,揭示生命的本质。目前研究大规模蛋白质相互作用的方法还主要是酵母双杂交及亲和层析,但研究者也发现,所得到的实验结果不能很好的匹配,仅有很少的重叠。这一方面说明蛋白质相互作用的复杂及多样性;另一方面也表明实验方法本身的不确定性及低重复性,对某些相互作用可能还无能为力。因此,对于大规模的蛋白质相互作用研究,不同方法的有机结合将给研究者提供更为丰富、准确的结果。
全文下载:http://www.bbioo.com/bbs/viewthread.php?tid=29316
4.参考文献
[1] Drewes G, Bouwmeester T. Global approaches to protein–protein
interactions. Curr Opin Cell Biol, 2003, 15(2): 199~205
[2] Bauer A, Kuster B. Affinity purification-mass spectrometry.Powerful tools
for the characterization of protein complexes.
[3]Eur J Biochem, 2003, 270(4): 570~578 [3] Fields S, Song O K. A novel
genetic system to detect protein- protein interactions. Nature, 1989,
340(6230): 245~246
[4] Uetz P, Giot L, Cagney G, et al. A comprehensive analysis of
protein-protein interactions in Saccharomyces cerevisiae. Nature, 2000,
403(6770): 623~627
[5] Ito T, Chiba T, Ozawa R, et al. A comprehensive two-hybrid analysis to
explore the yeast protein interactome. Proc Natl Acad Sci USA, 2001, 98(8):
4569~4574
[6] von Mering C, Krause R, Sne B, et al. Comparative assessment of
large-scale data sets of protein–protein interactions. Nature, 2002,
417(6887): 399~403
[7] Walhout A J, Temple G F, Brasch M A, et al. Gateway™ Recombinational
Cloning: Application to the cloning of large numbers of open reading frames or
ORFeomes. Methods Enzymol, 2000, 328: 575~592
[8] Rain J C, Selig L, De Reuse H, et al. The protein-protein interaction map
of Helicobacter pylori. Nature, 2001, 409 (6817): 211~215
[9] Giot L, Bader J S, Brouwer C, et al. A protein interaction map of
Drosophila melanogaster. Science, 2003, 302(5651): 1727~1736
[10] Li S M, Armstrong C M, Bertin N, et al. A map of the interactome network
of the metazoan C. elegans. Science, 2004, 303(5657): 540~543
[11] Stelzl U, Worm U, Lalowski M, et al. A human proteinprotein interaction
network: a resource for annotating the proteome. Cell, 2005, 122(6): 957~968
[12] Rual J F, Venkatesan K, Hao T, et al. Towards a proteomescale map of the
human protein–protein interaction network. Nature, 2005, 437(7062): 1173~1178
[13] Rigaut G, Shevchenko A, Rutz B, et al. A generic protein purification
method for protein complex characterization and proteome exploration. Nat
Biotechnol, 1999, 17(10): 1030~1032
[14] Bauer A, Kuster B. Affinity purification-mass spectrometry. Powerful
tools for the characterization of protein complexes. Eur J Biochem, 2003,
270(4): 570~578
[15] Dziembowski A, Seraphin B. Recent developments in the analysis of protein
complexes. FEBS Lett, 2004, 556(1-3): 1~6
[16] Gavin A C, Bösche M, Krause R, et al. Functional organization of the
yeast proteome by systematic analysis of protein complexes. Nature, 2002,
415(6868): 141~147
[17] Forler D, Kocher T, Rode M, et al. An efficient protein complex
purification method for functional proteomics in higher eukaryotes. Nat
Biotechnol, 2003, 21(1): 89~92
[18] Zhou D, Ren J X, Ryan T M, et al. Rapid tagging of endogenous mouse genes
by recombineering and ES cell complementation of tetraploid blastocysts.
Nucleic Acids Res, 2004, 32(16): e128
[19] Honey S, Schneider B L, Schieltz D M, et al. A novel multiple affinity
purification tag and its use in identification of proteins associated with a
cyclin-CDK complex. Nucleic Acids Res, 2001, 29(4): E24
[20] Ho Y, Gruhler A, Heilbut A, et al. Systematic identification of protein
complexes in Saccharomyces cerevisiae by mass spectrometry. Nature, 2002,
415(6868): 180~183
[21] Zhong J, Haynes P A, Zhang S, et al. Development of a system for the
study of protein-protein interactions in planta: characterization of a
TATA-box binding protein complex in Oryza sativa. J Proteome Res, 2003, 2(5):
514~522 [22] Blagoev B, Ong S E, Kratchmarova I, et al. Temporal analysis of
phosphotyrosine-dependent signaling networks by quantitative proteomics. Nat
Biotechnol, 2004, 22(9): 1139~1145
[23] Zhu H, Bilgin M, Bangham R, et al. Global analysis of protein activities
using proteome chips. Science, 2001, 293 (5537): 2101~2105
[24] Espejo A, Cote J, Bednarek A, et al. A protein-domain microarray
identifies novel protein–protein interactions. Biochem J, 2002, 367(Pt3):
697~702
[25] Hiller R, Laffer S, Harwanegg C, et al. Microarrayed allergen molecules:
diagnostic gatekeepers for allergy treatment. FASEB J, 2002, 16(3): 414~416
[26] Sreekumar A, Nyati M K, Varambally S, et al. Profiling of cancer cells
using protein microarrays: discovery of novel radiation-regulated proteins.
Cancer Res, 2001, 61(20): 7585~7593
[27] Ptacek J, Devgan G, Michaud G, et al. Global analysis of protein
phosphorylation in yeast. Nature, 2005, 438(7068): 679~684
-
综述
-
焦点事件

-
焦点事件
-
焦点事件












