E.Z.N.A.® Total RNA Midi Kit Protocol- DNase I digestion Protocol
实验概要
E.Z.N.A.® Total RNA Midiprep Kit provides a rapid and easy method for the isolation of up to 600 ug of total RNA from cultured eukaryotic cells, tissues, or bacteria. The kit allows single or multiple, simultaneous processing of samples in less than 40 min. Normally, 5 x 106 - 1 x 108 eukaryotic cells, 5 x 108-1 x 1010 bacterial cells, or 25- 200 mg tissue can be used in a single experiment. There is no need for phenol/chloroform extractions, and time-consuming steps such as CsCl gradient ultracentrifugation, and precipitation with isopropanol or LiCl, are eliminated. While this kit may be used for isolation of RNA from whole blood, we recommend you use the E.Z.N.A.® Blood RNA Kit (product # R6614/R6615) as it is specifically designed for effective hemolysis and hemoglobin removal and gives higher RNA yields.
RNA purified using the E.Z.N.A.® Total RNA method is ready for applications such as RT-PCR*, Northern blotting, poly A RNA (mRNA) purification, nuclease protection, and in vitro translation.
实验原理
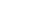
The E.Z.N.A. Total RNA Midiprep Kits use the reversible binding properties of HiBind® matrix, a new silica-based material. This is combined with the speed of mini-column spin technology. A specifically formulated high salt buffer system allows more than 600 ug of RNA molecules greater than 200 bases to bind to the matrix. Cells or tissues are first lysed under denaturing conditions that practically inactivate RNases. After the homogenization process by either bead-milling or rotor-stator homogenizer, samples are then applied to the HiBind® Midi spin columns to which total RNA binds, after few quick washing step, cellular debris and other contaminants are effectively washed away. High quality RNA is finally eluted in DEPC-treated sterile water.
实验步骤
Since HiBind® RNA resin and spin-column technology actually removes most of DNA without the DNase treatment, it is not necessary to do DNase digestion for most downstream applications. However, certain sensitive RNA applications might require further DNA removal. Following steps provide on-membrane DNase I digestion:( see DNase I cat.# E1091for detail information)
1. Follow the standard protocol until the samples completely pass through the HiBind RNA column (step1-4). Prepare the following:
1) For each HiBind® RNA column, prepare the DNase I digestion reaction mix as follows:
OBI DNase I Digestion Buffer 176 ul
RNase-free DNase I (20 Kunitz unites/ul) 4 ul
Total volume 180 ul
Note:
a. DNase I is very sensitive for physical denaturaion, so do not vortex this DNase I mixture. Mix gently by inverting the tube. Prepare the fresh DNase I digestion mixture before RNA isolation.
b. OBI DNase I digestion buffer is supplied with OBI RNase-free Dnase set.
c. Standard Dnase buffers are not compatible with on-membrane Dnase digestion.
2) Pipet 180 ul of the DNase I digestion reaction mix directly onto the surface of HiBind® RNA resin in each column. Make sure to pipet the Dnase I digestion mixture directly onto the membrane. Dnase I digestion will not be complete if some of the mix stick to the wall or the O-ring of the HiBind® RNA column.
3) Incubate at room temperature(25-30°C) for 15 minutes
2. Place column into a new 15 ml collection tube, and add 2 ml RNA Wash Buffer I. Place the column at benchtop for 5 minutes. Centrifuge at 4,000-5,000 x g for 5 minutes and discard flow-through. Reuse the collection tube in step 3.
3. Place column in the same 8ml collection tube, and add 2ml RNA Wash Buffer II diluted with ethanol. Centrifuge at 4,000-5,000 x g for 5 minutes and discard flow-through. Reuse the collection tube in step 4.
Note: Wash Buffer II Concentrate must be diluted with absolute ethanol before use. Refer to label on bottle for directions.
4. Wash column with a second 2 ml of Wash Buffer II as in step 3. Centrifuge and discard flow-through. Then with the collection tube empty, centrifuge the spin cartridge for 5 min at 5000 x g to completely dry the HiBind® matrix.
5. Elution of RNA. Transfer the column to a clean 15 ml microfuge tube (not supplied with kit) and elute the RNA with 250- 500 ul of DEPC-treated water (supplied with kit). Make sure to add water directly onto column matrix. Let it stand for 1 minute. Centrifuge 3 min at 8000x g to elute RNA. A second elution may be necessary if the expected yield of RNA >50 ug.
Alternatively, RNA may be eluted with a greater volume of water. While additional elutions increase total RNA yield, the concentration will be lowered since more than 80% of RNA is recovered with the first elution. Pre-heating the water to 70°C before adding to column and incubating column 5 min at room temperature before centrifugation may increase yields.
-
项目成果

-
焦点事件

-
焦点事件

-
科技前沿
-
科技前沿

-
科技前沿

-
科技前沿

-
技术原理
-
焦点事件

-
项目成果
-
产品技术

-
焦点事件

-
焦点事件

-
项目成果
-
焦点事件
-
项目成果

-
项目成果